Spotlight - JRG Banito
Sarcoma genetics and epigenetics
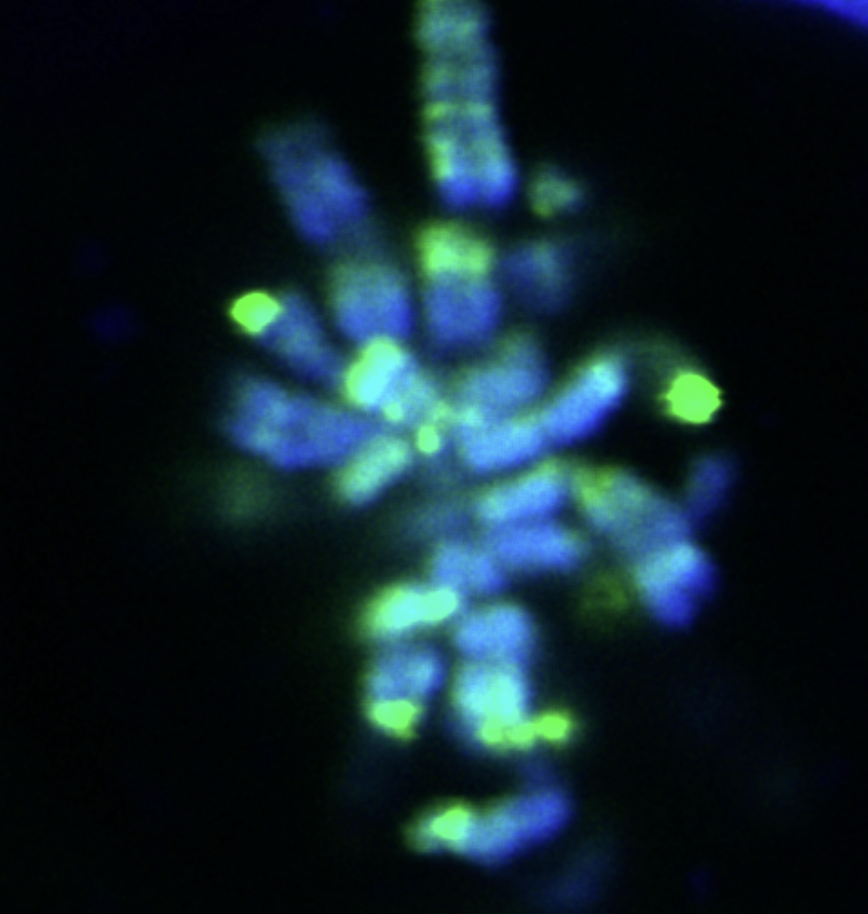
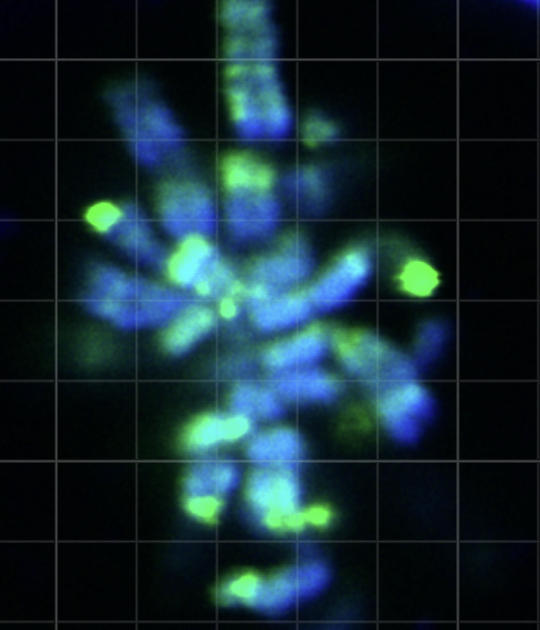
Gene fusions
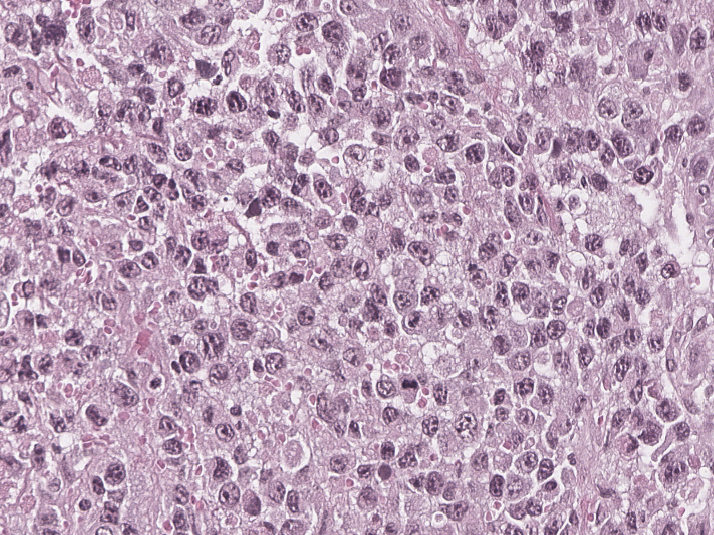
Sarcomas are often characterized by the presence of oncogenic gene fusions resulting from chromosomal translocations. By generating chimeric proteins that are not present in normal cells, they are rational drug targets. Still, how many of these genetic alterations drive tumorigenesis in sarcomas remains unclear. A recurrent theme in sarcoma biology, is the implication of genes encoding chromatin-modifying activities and transcription factors. We use a diverse set of assays to understand how these oncogenic alterations deregulate normal gene expression programs leading to cancer.
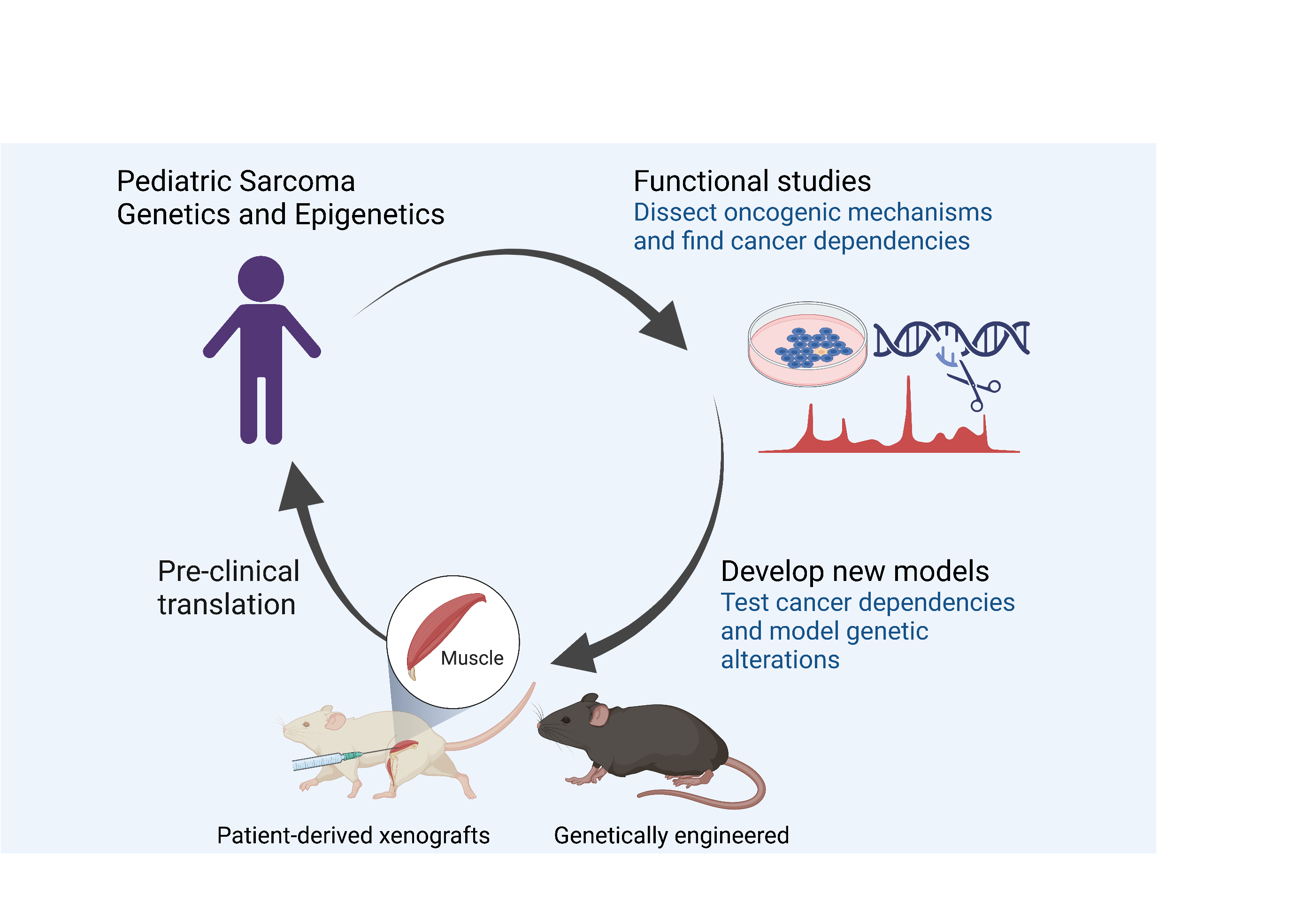
Model sarcoma genetic alterations in vivo
Sarcomas are an extremely heterogeneous group of tumors that can arise in a multitude of tissues and linages but their cells of origin remain elusive. More flexible and versatile models are in need to rapidly assess the impact of different genetic alterations in vivo. We established a somatic, fast and versatile, yet genetically controlled in vivo mouse modeling platform for soft-tissue sarcomas (EPO-GEMM/electroporation-based genetically-engineered mouse models). The method is based on a mix of transposon plasmids for oncogene delivery, and CRISPR/Cas9 plasmids expressing small guide RNAs (sgRNAs) directed against tumor suppressor genes, delivered by muscle electroporation. We are taking advantage of this flexible models to optimize immunotherapies in sarcoma and understand mechanisms of tumor maintenance by gene fusions in an immunocompetent background.
- Claudia Winter (Lab manager, TA)
- Dimtry Lupar (Colony manager, TA)
- Nezha Benabdallah, PhD (Postdoc)
- Roland Imle, MD (Postdoc)
- Vineet Dalal, PhD (Postdoc)
- Anastasija Pejkovska (PhD student)
- Afroditi Sotiriou (PhD student)
- Blösel, Daniel (PhD student)
- Clara Gade (MD student)
- Sara Placke (Master student)
- Felix Kommoss (Physician-Scientist)